Interdisciplinary project combining genomics, art, and graphic design to develop data visualization techniques making complex scientific data accessible to non-scientists. Co-PI with Chung-Jui Tsai (Warnell / Institute of Bioinformatics, UGA). All four NSF peer reviewers praised the science-art integration and design education component. Revision in progress.
Research Proposal
The Scientific Question
Aspen trees cover more of the earth's surface than almost any other tree species, yet they cannot interbreed with their closest relatives. Cottonwoods, balsam poplars, and aspens all belong to the genus _Populus_ and share strikingly similar genomes, but controlled crosses between aspen and these neighboring species consistently fail. The reason has been unknown for decades. Working from the haplotype-resolved aspen genome her lab produced (Zhou et al., 2023, _Plant Journal_, cover story) — one of the first genomic assemblies to accurately map highly repetitive sequences — Dr. Chung-Jui Tsai and her team identified a candidate explanation: roughly seven percent of the aspen genome consists of a single repeating DNA unit, called M147, organized in tandem arrays that stretch across millions of base pairs per chromosome. This sequence is absent in every other _Populus_ section, and the single exception — a tropical aspen that lacks M147 — is also the one aspen that can occasionally hybridize across sections. A second aspen-exclusive feature, a variant of CENH3 (a protein essential to chromosome function during cell division), co-occurs with M147 in a pattern too consistent to dismiss as coincidence. The proposed NSF research pursues the hypothesis that M147 arrays and CENH3.2 interact to enforce reproductive isolation — a finding that would reframe satellite DNA as a functional actor in species-level evolution and open new possibilities for _Populus_ breeding. The implications extend beyond trees: satellite DNA appears throughout eukaryotic genomes including human chromosomes, where its functions remain poorly understood.
Satellite DNA Art Studio
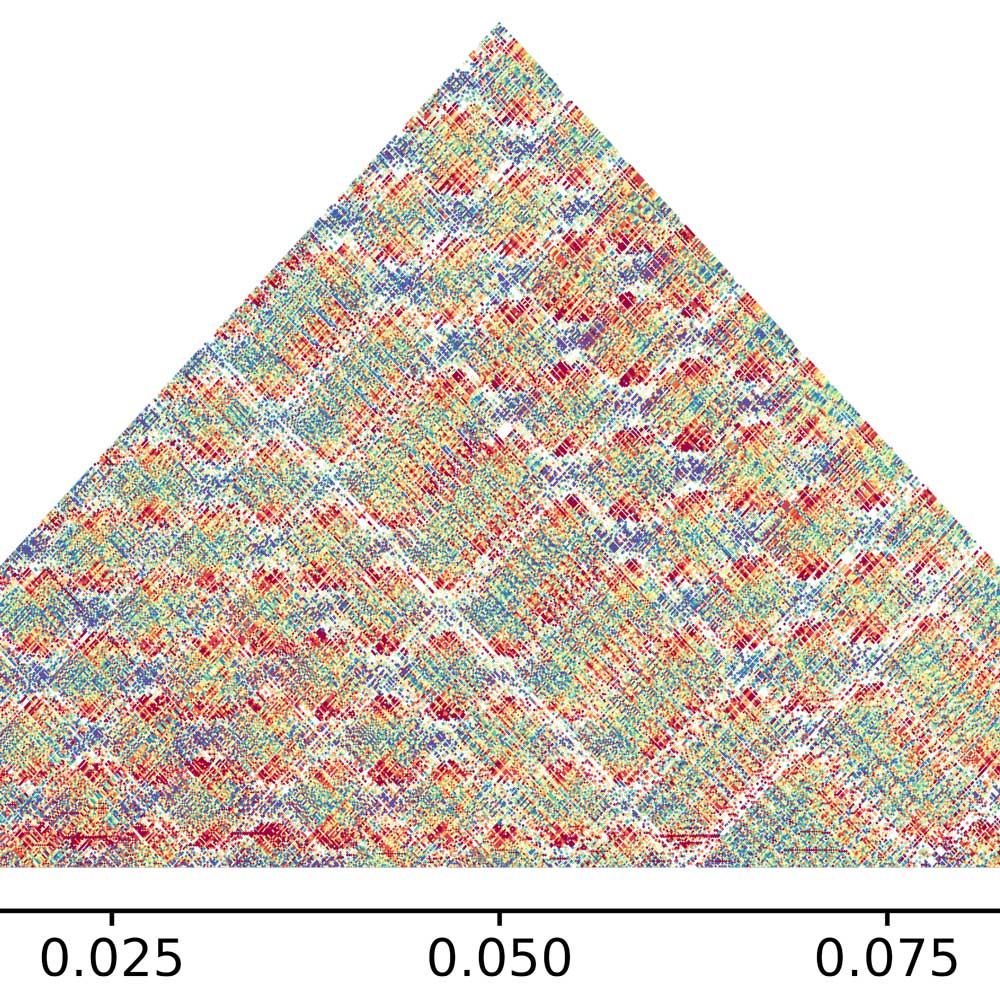
My role as co-PI centers on Activity 2 of the proposal's Broader Impacts plan — a component I designed and titled the _Satellite DNA Art Studio_. The scientific premise motivating this work is not merely that genomic data is difficult for non-specialists to access, but that the dominant modes of scientific data presentation — statistical tables, sequence alignment diagrams, heat maps — are themselves an epistemological problem. They are legible only to those already trained in their conventions, which means the most consequential genomic findings of the past decade have been effectively invisible to the public whose taxes funded them.
My research question within this collaboration is design-methodological: how can the principles and tools of graphic design be applied to genomic datasets not to make them decorative, but to make them genuinely intelligible — to translate structural complexity into visual form that communicates accurately to audiences outside the specialist community? This question sits at the intersection of communication design, data visualization, and the emerging field of science-art integration, and it is not rhetorical. Different visual representations of the same dataset produce measurably different comprehension outcomes. The problem of making M147 visible — literally: helping a viewer understand what it means that seven percent of a tree's DNA is a single repeating sequence — is a design problem with real epistemic stakes.
The proposed pedagogy integrates this research question directly into the graphic design curriculum. Under my supervision, students in Graphic Systems (ARGD 3020) — an upper-level course focused on visual problem-solving for complex, abstract content — would work with actual genomic datasets from the Tsai lab, developing original visualization approaches ranging from graphic simplification and information design to mixed media, three-dimensional installation, and generative AI. The student work functions not as illustration of pre-existing conclusions, but as a research catalog: multiple, competing visual translations of the same data, each representing a different design strategy, each legible (or not) to different audiences. That catalog is itself the scholarly output — a documented investigation of which visual approaches produce comprehension, and for whom.
A dedicated exhibition pipeline was designed to move the work into public space: campus galleries at the Lamar Dodd School of Art, the Georgia Museum of Art, public libraries, parks, and outdoor venues in Athens, with national exhibition as a longer-term trajectory. A public-facing website would serve as a permanent online documentation of both the science and the design process.
To develop the pilot version of this curriculum in advance of full funding, I was awarded a $1,000 mini-grant from the UGA Arts Collaborative. This grant is supporting the design of an intensive 2–3 day cross-disciplinary workshop, bringing together students from graphic design and the biological sciences to work collaboratively with satellite DNA data. The workshop is in active planning and represents the first operational phase of a curriculum that, at full scale, would run across a semester and produce an exhibitable body of work.
Peer Review Record
The NSF proposal was reviewed by five external specialists and received a panel rating of Very Good across four of five reviews. All five reviewers addressed the Broader Impacts component explicitly; none raised criticisms. Two reviewers quoted the science-art integration as a specific strength for the Broader Impacts criteria. One wrote that "the team assembled to provide the teaching and training associated with this project in both biological sciences and the arts is compelling and likely to generate interest to students and the public." Another called working with a graphic design specialist "a very nice touch" and noted it as a genuine opportunity for graphic design students. A third described the activities as "well-considered" and "creative," specifically naming the inclusion of art students as a novel approach to public engagement.
The only substantive request across all reviews was the addition of audience metrics for future submissions — number of non-science majors reached, exhibition visitors, online gallery engagement. This is an addressable note that I am building into the documentation framework for the workshop and any subsequent curriculum implementation.
The scientific component was not funded due to insufficient preliminary data on the CENH3.2 hypothesis — a genetics question entirely outside my scope. My component received no criticism and the peer review record is on file.
---
Status and Trajectory
The grant is currently under revision. Dr. Tsai and Senior Research Associate Ran Zhou are leading the scientific revision; resubmission timeline is pending. My Broader Impacts design — the curriculum framework, exhibition pipeline, and pedagogy — is fully developed and independent of the grant timeline. The pilot workshop supported by the Arts Collaborative mini-grant proceeds in parallel, generating preliminary documentation and pedagogical data that will strengthen the next submission. When the grant is resubmitted, my component will include the audience metrics requested by reviewers and the documented outcomes of the pilot workshop as evidence of feasibility.
This project represents a methodological strand that runs across my research program: the conviction that design is not downstream of knowledge production, but constitutive of it. How data is represented determines what can be seen and understood. The _Satellite DNA Art Studio_ applies that conviction to one of the most technically inaccessible datasets in contemporary biology — and asks graphic design students to treat scientific incomprehensibility not as a limitation of the audience, but as a design problem waiting to be solved.
Data Visualization Pilot Workshop
The pilot workshop is the first operational phase of the _Satellite DNA Art Studio_, the Broader Impacts component of the NSF _Function and Dynamics of Aspen-Specific M147 Tandem Repeat Arrays_ proposal. Its purpose is to test, document, and refine a condensed version of the cross-disciplinary curriculum that Kappenstein is developing as co-PI, generating both pedagogical data and preliminary evidence of feasibility for the NSF revision.
The workshop brings together a small cohort of 6–10 students from graphic design and the biological sciences for three days of intensive, collaborative work with actual genomic datasets provided by the Tsai lab. It is structured as a research instrument as much as a pedagogical event: the animating question is not "can students make science visually appealing?" but "which design strategies produce genuine comprehension of complex genomic data in non-specialist audiences?" Each day advances that question in a different register.
Day 1 establishes shared ground. Biology and graphic design students encounter the same dataset — chromosome maps, M147 distribution diagrams, sequence abundance visualizations — from their respective disciplinary positions. A structured looking exercise makes the gap between scientific literacy and visual legibility explicit and productive: biology students articulate what the data shows; design students articulate what it communicates, and what it fails to communicate. That gap is the brief. Each participant or pair selects one aspect of the M147 dataset and a defined non-specialist audience, and produces first-concept sketches by end of day.
Day 2 is design development and structured critique. Participants work in their chosen medium — print, mixed media, digital, three-dimensional — without format constraint, translating their concept into visual form. A mid-day critique crosses the disciplinary divide deliberately: biology students evaluate scientific accuracy; graphic design students evaluate communicability. Points of disagreement between the two are treated as data, not error. Participants iterate based on the critique, and end the day with near-final work documented and brief written reflections collected. Those reflections constitute the first layer of pedagogical documentation for the NSF revision.
Day 3 moves to installation and public presentation. Work is installed as prototypes in the gallery space at the Lamar Dodd School of Art — resolved enough to exhibit, explicitly framed as first-iteration research rather than finished product. A small invited audience attends a brief public presentation in which participants explain their work and their design decisions. Professional documentation of the installed work is produced. A structured debrief closes the workshop, generating the facilitation notes, audience metrics, and participant reflection data that will form the evidence base for the full-scale curriculum proposal.
The $1,000 Arts Collaborative mini-grant supports materials, installation hardware, catering across the three working days, and documentation. All facilitation is provided by Kappenstein as in-kind contribution; data, datasets, and scientific expertise are provided by Dr. Tsai and Senior Research Associate Ran Zhou as in-kind from the Warnell School. Both contributions are being documented explicitly as evidence of institutional commitment for the NSF resubmission.
The workshop is currently in active planning. Its outputs — exhibited prototypes, participant reflections, audience metrics, and a documented exhibition record — will address the one substantive request made by NSF reviewers across all five reviews: the addition of measurable audience impact data to the Broader Impacts component. No reviewer criticized the _Satellite DNA Art Studio_ concept; two cited it as a specific strength. The pilot workshop transforms that concept into evidence.